Welcome to CLEP’s documentation!¶
Release notes : https://github.com/hybrid-kg/clep/releases

CLEP: A Hybrid Data- and Knowledge- Driven Framework for Generating Patient Representations.¶
CLEP has three main subgroups: sample_scoring, embedding, classify.
The
sample_scoringmodule generates a score for every patient-feature pair.
2. The embedding module overlays the patients on the prior knowledge in-order generate a new KG, whose embedding
is generated using KGE models from PyKEEN(Ali, et al.,2020).
3. The classify module classifies the generated embedding model (or any data that is passed to it) using generic
classification models.
General info¶
CLEP is a framework that contains novel methods for generating patient representations from any patient level data and its corresponding prior knowledge encoded in a knowledge graph. The framework is depicted in the graphic below
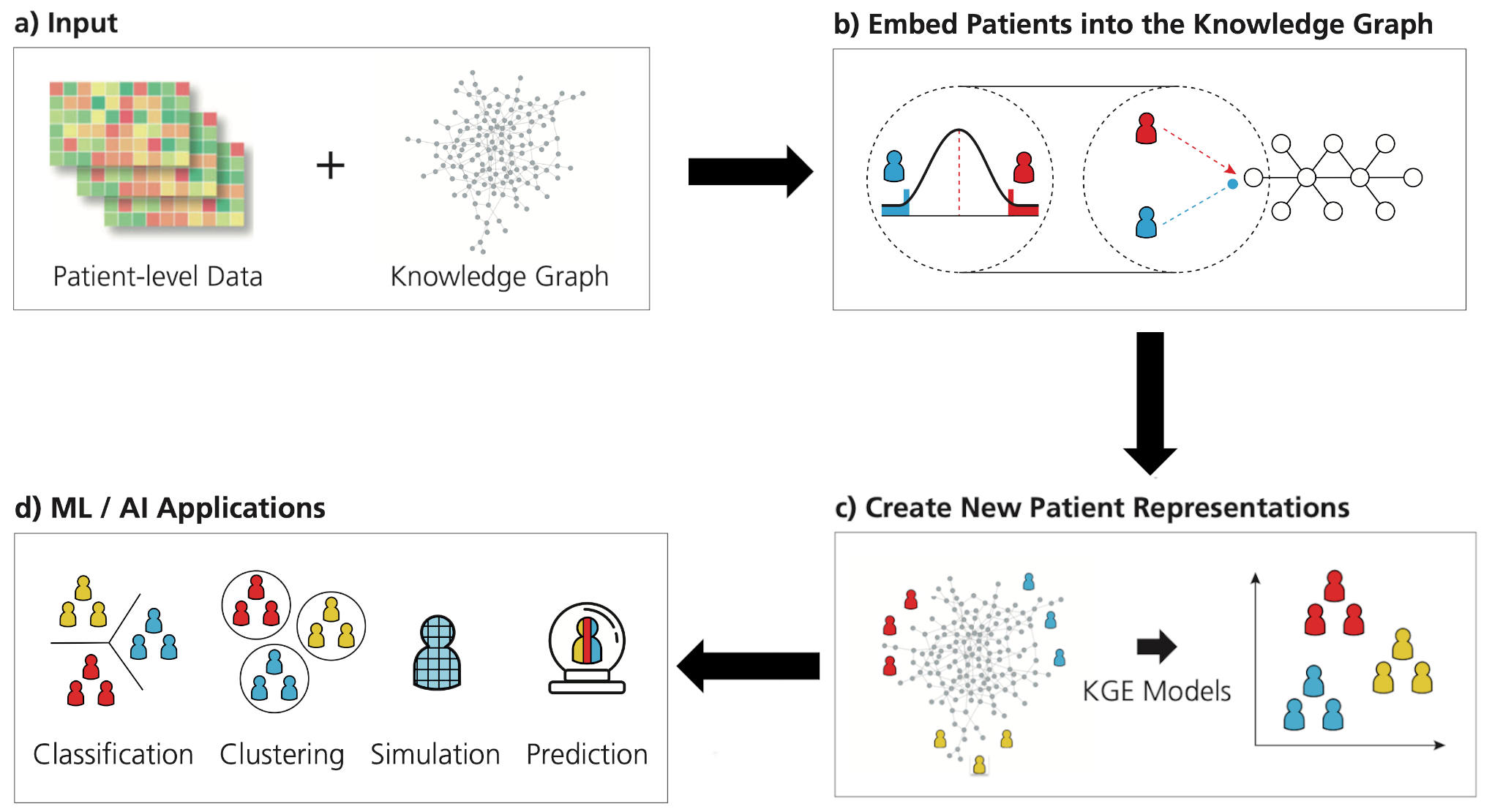
Installation¶
In-order to install CLEP, an installation of R is required with a copy of Limma. Once they are installed, you can install CLEP package from pypi.
# Use pip to install the latest release
$ python3 -m pip install clep
You may instead want to use the development version from Github, by running
$ python3 -m pip install git+https://github.com/hybrid-kg/clep.git
For contributors, the repository can be cloned from GitHub and installed in editable mode using:
$ git clone https://github.com/hybrid-kg/clep.git
$ cd clep
$ python3 -m pip install -e .
Dependency¶
Python 3.6+
Installation of R
Mandatory¶
Numpy
Scipy
Pandas
Matplotlib
rpy2 (for limma)
Limma package from bioconductor
For API information to use this library, see the Developmental Guide.
Disclaimer¶
CLEP is a scientific software that has been developed in an academic capacity, and thus comes with no warranty or guarantee of maintenance, support, or back-up of data.
